What's new?
Dissecting metabolism by heat measurements
metabolic heat production during bacterial growth
Source: Prof. Dr. Fahmy, Karim
Metabolic heat released upon biochemical reactions is an extremely sensitive sum signal that can be recorded from cultured cells and living multicellular organisms. Although it is not specific in itself, it carries important information on the metabolic state and biochemical efficiency of metabolism.
We use isothermal microcalorimetry to study the growth of microorganisms to reveal metal toxicity and adaptational strategies to such stressors. A hallmark of our recent research is the conservation of relatively simple patterns of heat release from bacteria, eukaryotic cell cultures, and plant cells. Nutrient dependence of microbial metabolic activity as previously formulated in the famous Monod equation for the exponential growth phase of bacteria only appear to hold throughout the entire life span of a bacterial culture. This discovery has been made possible by the development of a mathematical approach that describes growth metabolism relations in the enthalpy domain, rather than the conventionally used time domain. Current activities aim at automizing these evaluations and develop a web tool to evaluate thermograms from a variety of organisms.
Excellence Cluster "Physics of Life" of the TU-Dresden and its partners
The Biophysics Department received funding from the Helmholtz Society for the participation in the Excellence Cluster "Physics of Life". We contribute our expertise in vibrational spectroscopy using static and time-resolved Fourier Transform spectroscopy. Research on water structures in "crowded" protein environments is performed in collaboration with the Institute of Radiation Physics where Tera-Hertz pump optical propbe techniques are being established as a part of the "Biophysics beamline" at the research infrastructure TELBE. The cluster aims at elucidating the physical principles that govern the dynamic organization of living matter. On the molecular scale, the focus lies on the self-organization of proteins in liquid phases within the cytoplasm. The user facility TELBE at the HZDR provides pulsed THz sources with high repetition and intensity. This enables unique studies of the dielectric dynamics in such phases and offers extraordinary synergies with the complementary experimental approaches by our partners.
DEBs: DNA-encircled lipid bilayers
In a joint project with the TU-Dresden and the Rockefeller University New York, USA, we have recently succeeded in producing DNA-encircled lipid bilayers. Their size is similar to nanodiscs formed by membrane-scaffolding proteins. In contrast to protein-based scaffolds with limited size variability, we show that dsDNA minicircles whose size can be precisely controlled enable lipid bilayer formation in a self-assembly process. At the same time, hybridization-based approaches will allow linking such units to substrates and other DNA structures. Thus, The technology enables new experimental systems for biophysical membrane research. For more information see Katarina Iric et al. Nanoscale 93(2018)
Synthesis of DEBs. (a) Short oligonucleotides with two or four phosphorothioates are alkylated with alkyl-iodides (red). (b) A circular single-stranded template is synthesized by enzymatic splint ligation from a long, linear oligonucleotide (grey). Residual splints (green) and linear templates are digested by exonuclease treatment. (c) A denaturing PAGE gel confirms the synthesis of the single-stranded minicircle (ssMC). M, molecular size marker (nt); lane 1, linear long oligonucleotide; lane 2, ligation reaction before exonuclease treatment; lane 3, exonuclease digest. (d) The ssMC is hybridized with seven alkylated oligonucleotides into double-stranded MCs (dsMC). The position of alkylations, and the intrinsic curvature of the A-tracts (grey line in the center of the helix, exaggerated) in a model of an A-tract32,33 (adapted from PDB structure 1FZX). Adenosines are coloured red, thymidines blue. (e) Native PAGE gel. M, marker (base pairs); 1, linear long oligonucleotide; 2, the assembled dsMC complex. (f ) The dsMC is incubated with phospholipids (blue) to form a mature DEB (g).
(a) tSEM image of a DEB with 28 decyl modifications. (b) Elution profile of a size exclusion chromatography (SEC) run of a DEB preparation. Fraction 1 contains dimeric DEBs (h), fraction 2 monomers (f ). Scale bars, 50 nm.
The Department of Biophysics hosted the Thermal Activity Monitor (TAM) User Meeting 2017 on Microcalorimetry on Oct. 11th, 2017
MDR-Sachsenspiegel Television News on Radioecological Research on uranium toxicity
We have organized and enjoyed an exciting "Joint Meeting of Czech and German Biophysicists 2017"
run by the Section "Molecular Biophysics" of the German Biophysical Society
A single particle approach to studying membrane protein structure and function
The insertion of a single metal-transporting membrane protein into a nanodisc (image) has allowed studying a fundamental ques
tion in metal-transport across biological membranes: How is the hydration of the mostly hydrophobic interior of a membrane protein controlled by the purely hydrophobic interactions between the protein and its surrounding lipidic phase?
Using the copper-transporting bacterial ATPase CopA from Legionella pneumophila, we have exploited the accessibility of the native cysteine residues of its metal-binding site for chemical modification with a flurophore. The latter reacts strongly to the polarity and mobility of the local dipoles in its viccinity. Thereby, it was possible to monitor the dipole relaxation after optical excitation of the chromophore within specific sites inside the membrane protein. The analysis of time-dependent shifts of the fluorescence emission maximum revealed that the membrane lateral pressure has a remarkable influence on intra-membrane protein hydration and is thus likely to support transient hydration / dehydration events that occur during the cyclic ATP-driven metal transport in this class of P1B-type ATPases. The results of this study have been published in Fischermeier et al., Angewandte Chemie Int. Ed., 2017, 56: pp 1269-1272.
The use of the depicted self-assembly of a membrane-scaffolding protein (green) around a lipid bilayer that harbours the membrane protein has not only helped performing optical spectroscopy.
It is also a highly attractive system for future use in single particle diffraction experiments that will be enabled by new x-ray free electron lasers such as the European XFEL in Hamburg.
Paramagnetic DNA Nanostructures
DNA-origami are nanostructures that are formed by the rational design of DNA scaffold and staple strands, using base complementarity to fold the scaffold into a predicted arbitrary shape. We have studied the potential of paramagnetic decoration to confer magnetic properties to such nanostructures. We show that Eu3+, a paramagnetic lanthanide, is strongly coordinated by two differently shaped DNA nanostructures. Whereas natural DNA double strands would change their geometry and aggregate upon neutralization of heir surface charge by Eu3+, the DNA-origami exhibit sufficient rigidity to preserve their structure even when their molecular surfaces are are close to saturated with the trivalent lanthanide. This allows producing DNA-based structures with magnetic properties based on the paramagnetism of a trivalent lanthanide. Lars Opherden et al., Langmuir, 2014, 30: pp 8152–8159.
Figure 1 (left): DNA origami triangles(left AFM image) preserve their shape when they bind large amounts of Eu3+ (about 2000 per DNA origami) before they aggregate (obvious from the sudden decrease of their UV-vis absorption, red curve). The shape of the DNA-origami is preserved even under conditions of excess Eu3+ (right AFM image). (One arm of the triangle is ~130 nm long). (Move mouse over image to enlarge.)
Figure 2 (right): Eu3+ affects the magnetic circular dichroism (MCD) of DNA origami in a superstructure-dependent manner. The replacement of the natural Mg2+ counterion to the DNA phosphate backbone by Eu3+ strongly affects the MCD of DNA origami triangles below 225 nm, whereas in six-helical bundles, only the weak change between 280 and 300 nm is observed. The Eu3+ effect on the MCD of of both superstructures differs from that of on genomic DNA, evidencing a superstructure-dependent interaction of the lanthanide with the DNA structures. In addition, time-resolved luminescence of Eu3+ bound to the DNA origami supports the conclusion that the coordination geometry of Eu3+ is different in the two superstructures and exhibits an asymmetric coordination environment. Such binding site asymmetries form the basis for exploiting the paramagnetism of the lanthanide to orient DNA-based nanostructures in external magnetic fields. (Move mouse over image to enlarge.)
Calorimetric monitoring of metal toxicity in organisms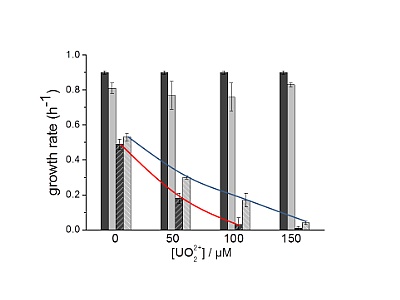
Both prokaryotic and eukaryotic organisms possess mechanisms for the detoxification of heavy metals, which are found among distantly related species. We investigated the role of intracellular glutathione (GSH), which in a large number of taxa plays a role in the protection against the toxicity of common heavy metals. Using a Lactococcus lactistrain with an inducible GSH synthesis pathway it was possible to measure the influence of GSH on the toxicity of uranium. By microcalorimetric measurements of the metabolic heat during cultivation, it was shown that intracellular GSH attenuates the toxicity of uranium at a concentration in the range of 10-150 µM. The toxicity was derived from the effect on the growth rate during late exponential growth. The rate could be accurately determined from the metabolic heat production of the culture.
This is the first demonstration of an intracellular role of GSH in uranyl toxicity independent of interference from oxidative stress because the cukture were grown anaerobically. Isothermal titration calorimetry further revealed the endothermic binding of U(VI) to the carboxyl group(s) of GSH, rather than to the reducing thiol group involved in copper interactions. The data indicate that the primary detoxifying mechanism is the intracellular
sequestration of carboxyl-coordinated U(VI) into an insoluble complex with GSH. Obeid et al. Appl Environ Microbiol., 2016, 82:3563-3571.
Structure and Energetics of Europium Polycarboxylate Complexes
The mobility of actinides and other metal ions in the environment is largely influenced by their interaction with humic acids. The latter bind multivalent cations at their multiple carboxylic functions. We have studied the binding of the lanthanide Eu3+, a non-radioactive model for the physico-chemical behaviour of trivalent actinides, to benzenetetracarboxylic acid (BTC), a polycarboxylate model for humic acids. The figure shows the heat uptake during Eu BTC complex formation measured by isothermal titration calorimetry upon injection of BTC into a 1 mM Eu3+ solution. The integrated heats (lower panel) of the fast (red) and slow (blue) calorimetric response were evaluated separately and assigned to a 1:1 complex formation and an ensuing metal-mediated polymerization, respectively. Both processes are entropy-driven with ΔH of 14-17 kJ mol-1 and ΔS of 130-160 J mol-1 K-1. In combination with infrared, fluorescence spectroscopy and DFT calculations an energetically and structurally consistent model of the lanthanide-BTC interaction and polymerization (reaction scheme) has been derived (Barkleit et al. 2011).
Infrared spectroscopy reveals the physical basis of desiccation tolerance in an anhydrobiotic organism.
We have used FTIR-difference spectroscopy to study the effect of desiccation stress on the physical state of C.elegans larvae that are either competent or defficient of trehalose synthesis. The spectra indicate a critical role of trehalose on the reversibility of dessication-induced packing changes in lipidic components of the organism. The vibrational frequency of the CH2 stretching modes (dominated by lipidic acyl chains) change during desiccation of the live organism. Only in the presence of trehalose, these changes are reversed by subsequent rehydration. The physiological, morphological and spectroscopic traits of different C.elegans strains are well correlated (Erkut et al. CurrBiol 2011; Erkut et al. Worm 2012).
Picture (right): Infrared absorption spectra in the range of CH2 stretching modes of live C.elegans Dauer larvae. Top to bottom: initial IR absorption spectrum; absorption changes upon rehydration of dessicated larvae; absorption changes during dessication of larvae; absorption changes during dessication of extracted C.elegans lipids. Irreversible changes in hdyration / dehydration spectra of this trehalose defficient strain can be seen at 2858 and 2916 cm-1. (Move mouse over image to enlarge.)
Aqueous coordination chemistry and photochemistry of uranyl(VI) oxalate.
The bioavailability of uranyl in contaminated soils and waters depends on its solubility. The latter is strongly affacted by the uranyl redox state as U(VI) complexes exhibit high solubility as compared to the essentially unsoluble reduced U(IV) state. Light can additionally affect the redox state through photochemical reactions in uranyl complexes with organic ligands as in light exposed soils or plant tissues of contaminated areas. Using density functional theory (DFT) calculations, we revisited a classical problem of uranyl(VI) oxalate photochemical decomposition. Photoreactivities of uranyl(VI) oxalate complexes are found to correlate largely with ligand-structural arrangements. Importantly, the intramolecular photochemical reaction is inhibited when oxalate is bound to uranium exclusively in chelate binding mode. Previously proposed mechanisms involving a UO2(C2O4)22− (1:2) complex as the main photoreactive species are thus unlikely to apply, because the two oxalic acids are bound to uranium in a chelating binding mode. Our DFT results suggest that the relevant photoreactive species are UO2(C2O4)34− (1:3) and (UO2)2(C2O4)56− (2:5) complexes binding uranium in an unidentate fashion. These species go through decarboxylation upon excitation to the triplet state, which ensues the release of CO2 and reduction of U(VI) to the U(V) state (Tsushima et al., 2010).
Picture (right): DFT-result for the ground state geometry (bond lengths in Angstrom) of a uranyl:oxalate 2:5 complex. The oxalates in unidentate coordination (lower left and upper right) undergo decarboxylation upon photon-induced electron transfer and U(V) formation in the excited state of the actinide. (Move mouse over image to enlarge.)

